Protein Name: Dysbindin
|
UniprotKB/SwissProt ID: DTBP1_HUMAN (Q96EV8)
Gene Name: DTNBP1
Synonyms: BLOC1S8
Organism: Homo sapiens (Human).
Function: Component of the BLOC-1 complex, a complex that is required for normal biogenesis of lysosome-related organelles (LRO), such as platelet dense granules and melanosomes. In concert with the AP-3 complex, the BLOC-1 complex is required to target membrane protein cargos into vesicles assembled at cell bodies for delivery into neurites and nerve terminals. The BLOC-1 complex, in association with SNARE proteins, is also proposed to be involved in neurite extension. Associates with the BLOC-2 complex to facilitate the transport of TYRP1 independent of AP-3 function. Plays a role in synaptic vesicle trafficking and in neurotransmitter release. Plays a role in the regulation of cell surface exposure of DRD2. May play a role in actin cytoskeleton reorganization and neurite outgrowth. May modulate MAPK8 phosphorylation. Appears to promote neuronal transmission and viability through regulating the expression of SNAP25 and SYN1, modulating PI3-kinase-Akt signaling and influencing glutamatergic release. Regulates the expression of SYN1 through binding to its promoter. Modulates prefrontal cortical activity via the dopamine/D2 pathway.
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization: Isoform 1: Cytoplasm. Cytoplasmic vesicle membrane; Peripheral membrane protein; Cytoplasmic side. Endosome membrane; Peripheral membrane protein; Cytoplasmic side. Melanosome membrane; Peripheral membrane protein; Cytoplasmic side. Cell junction, synaps
| |
|
Protein disease:
|
| Disease database |
Database Entry |
Disease information | | KEGG | H00166 | Hermansky-Pudlak syndrome (HPS) | | OMIM | 181500 | SCHIZOPHRENIA; SCZD ;;SCHIZOAFFECTIVE DISORDER, INCLUDED | | OMIM | 607145 | DYSTROBREVIN-BINDING PROTEIN 1; DTNBP1 ;;DYSBINDIN;; SANDY, MOUSE, HOMOLOG OF; SDY | | OMIM | 614076 | HERMANSKY-PUDLAK SYNDROME 7; HPS7 | | HPRD | 06190 | Hermansky-pudlak syndrome 7 |
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 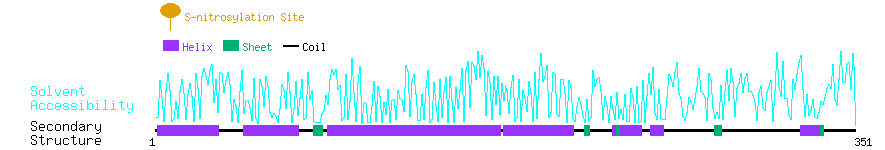 |
|

