Protein Name: Persulfide dioxygenase ETHE1, mitochondrial
|
UniprotKB/SwissProt ID: ETHE1_MOUSE (Q9DCM0)
Gene Name: Ethe1
Synonyms: Hsco
Organism: Mus musculus (Mouse).
Function: First described as a protein that can shuttle between the nucleus and the cytoplasm and suppress p53-induced apoptosis by sequestering the transcription factor RELA/NFKB3 in the cytoplasm and preventing its accumulation in the nucleus (By similarity). Sulfur dioxygenase that plays an essential role in hydrogen sulfide catabolism in the mitochondrial matrix. Hydrogen sulfide (H(2)S) is first oxidized by SQRDL, giving rise to cysteine persulfide residues. ETHE1 consumes molecular oxygen to catalyze the oxidation of the persulfide, once it has been transferred to a thiophilic acceptor, such as glutathione (R-SSH). Plays an important role in metabolic homeostasis in mitochondria by metabolizing hydrogen sulfide and preventing the accumulation of supraphysiological H(2)S levels that have toxic effects, due to the inhibition of cytochrome c oxidase.
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization: Cytoplasm (By similarity). Nucleus (By similarity). Mitochondrion matrix (By similarity).
| |
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 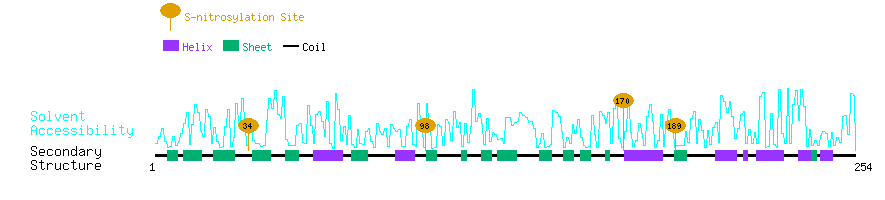 |
|

