Protein Name: 26S proteasome non-ATPase regulatory subunit 14
|
UniprotKB/SwissProt ID: PSDE_MOUSE (O35593)
Gene Name: Psmd14
Synonyms: Pad1
Organism: Mus musculus (Mouse).
Function: Metalloprotease component of the 26S proteasome that specifically cleaves 'Lys-63'-linked polyubiquitin chains. The 26S proteasome is involved in the ATP-dependent degradation of ubiquitinated proteins. Plays a role in response to double-strand breaks (DSBs): acts as a regulator of non-homologous end joining (NHEJ) by cleaving 'Lys-63'-linked polyubiquitin, thereby promoting retention of JMJD2A/KDM4A on chromatin and restricting TP53BP1 accumulation. Also involved in homologous recombination repair by promoting RAD51 loading (By similarity).
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization:
| |
|
Network with metabolic pathway:
|
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 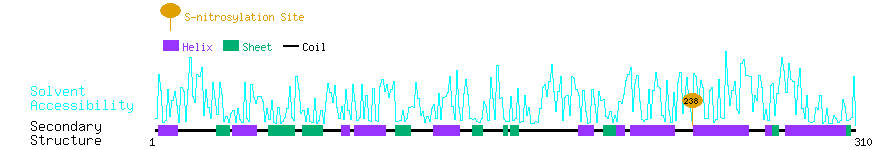 |
|

