Protein Name: Ubiquitin-conjugating enzyme E2 N
|
UniprotKB/SwissProt ID: UBE2N_MOUSE (P61089)
Gene Name: Ube2n
Synonyms: Blu
Organism: Mus musculus (Mouse).
Function: The UBE2V1-UBE2N and UBE2V2-UBE2N heterodimers catalyze the synthesis of non-canonical 'Lys-63'-linked polyubiquitin chains. This type of polyubiquitination does not lead to protein degradation by the proteasome. Mediates transcriptional activation of target genes. Plays a role in the control of progress through the cell cycle and differentiation. Plays a role in the error-free DNA repair pathway and contributes to the survival of cells after DNA damage. Acts together with the E3 ligases, HLTF and SHPRH, in the 'Lys-63'-linked poly-ubiquitination of PCNA upon genotoxic stress, which is required for DNA repair. Appears to act together with E3 ligase RNF5 in the 'Lys-63'-linked polyubiquitination of JKAMP thereby regulating JKAMP function by decreasing its association with components of the proteasome and ERAD (By similarity). Promotes TRIM5 capsid-specific restriction activity and the UBE2V1-UBE2N heterodimer acts in concert with TRIM5 to generate 'Lys-63'-linked polyubiquitin chains which activate the MAP3K7/TAK1 complex which in turn results in the induction and expression of NF-kappa-B and MAPK-responsive inflammatory genes (By similarity).
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization: Nucleus (By similarity). Cytoplasm (By similarity).
| |
|
Network with metabolic pathway:
|
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 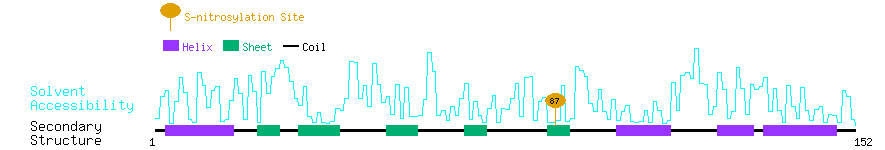 |
|

