Protein Name: Histone deacetylase 6
|
UniprotKB/SwissProt ID: HDAC6_MOUSE (Q9Z2V5)
Gene Name: Hdac6
Organism: Mus musculus (Mouse).
Function: Responsible for the deacetylation of lysine residues on the N-terminal part of the core histones (H2A, H2B, H3 and H4). Histone deacetylation gives a tag for epigenetic repression and plays an important role in transcriptional regulation, cell cycle progression and developmental events. Histone deacetylases act via the formation of large multiprotein complexes (By similarity). Plays a central role in microtubule-dependent cell motility via deacetylation of tubulin. In addition to its protein deacetylase activity, plays a key role in the degradation of misfolded proteins: when misfolded proteins are too abundant to be degraded by the chaperone refolding system and the ubiquitin-proteasome, mediates the transport of misfolded proteins to a cytoplasmic juxtanuclear structure called aggresome. Probably acts as an adapter that recognizes polyubiquitinated misfolded proteins and target them to the aggresome, facilitating their clearance by autophagy (By similarity).
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization: Nucleus. Cytoplasm. Note=It is mainly cytoplasmic, where it is associated with microtubules.
| |
|
Network with metabolic pathway:
|
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 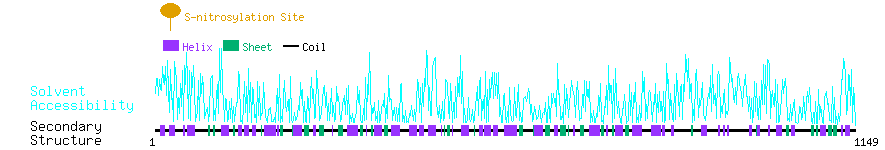 |
|

