Protein Name: Nitric oxide synthase, endothelial
|
UniprotKB/SwissProt ID: NOS3_HUMAN (P29474)
Gene Name: NOS3
Organism: Homo sapiens (Human).
Function: Produces nitric oxide (NO) which is implicated in vascular smooth muscle relaxation through a cGMP-mediated signal transduction pathway. NO mediates vascular endothelial growth factor (VEGF)-induced angiogenesis in coronary vessels and promotes blood clotting through the activation of platelets. Isoform eNOS13C: Lacks eNOS activity, dominant-negative form that may down-regulate eNOS activity by forming heterodimers with isoform 1.
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization: Cell membrane. Membrane, caveola. Cytoplasm, cytoskeleton. Golgi apparatus. Note=Specifically associates with actin cytoskeleton in the G2 phase of the cell cycle; which is favored by interaction with NOSIP and results in a reduced enzymatic activity.
|
PDB :
( If your security settings prevent Jmol from running, please register http://140.138.144.145/ as a safe location in your Java settings. ) |
|
Protein disease:
|
| Disease database |
Database Entry |
Disease information | | OMIM | 104300 | ALZHEIMER DISEASE; AD ;;PRESENILE AND SENILE DEMENTIA ALZHEIMER DISEASE, FAMILIAL, 1, INCLUD | | OMIM | 145500 | HYPERTENSION, ESSENTIAL ;;EHT | | OMIM | 163729 | NITRIC OXIDE SYNTHASE 3; NOS3 ;;NITRIC OXIDE SYNTHASE, ENDOTHELIAL; ENOS CORONARY ARTERY SPA | | OMIM | 601367 | STROKE, ISCHEMIC ;;CEREBROVASCULAR ACCIDENT;; CEREBRAL INFARCTION | | HPRD | 01224 | Coronary spasms, susceptibility to |
|
Network with metabolic pathway:
|
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 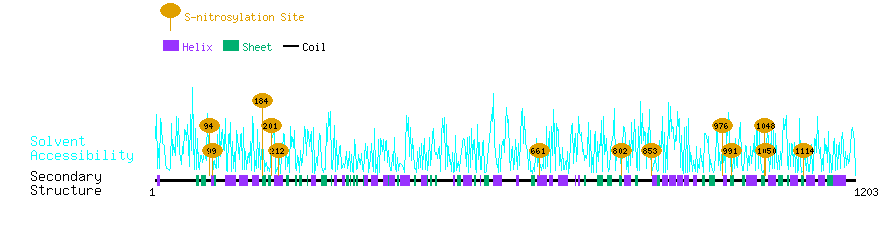 |
|
