Protein Name: NAD-dependent protein deacylase sirtuin-5, mitochondrial
|
UniprotKB/SwissProt ID: SIR5_MOUSE (Q8K2C6)
Gene Name: Sirt5
Synonyms: Sir2l5
Organism: Mus musculus (Mouse).
Function: NAD-dependent lysine demalonylase and desuccinylase that specifically removes malonyl and succinyl groups on target proteins. Activates CPS1 and contributes to the regulation of blood ammonia levels during prolonged fasting: acts by mediating desuccinylation of CPS1, thereby increasing CPS1 activity in response to elevated NAD levels during fasting. Activates SOD1 by mediating its desuccinylation, leading to reduced reactive oxygen species. Has weak NAD-dependent protein deacetylase activity; however this activity may not be physiologically relevant in vivo. Can deacetylate cytochrome c (CYCS) and a number of other proteins in vitro such as Uox.
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization: Mitochondrion. Cytoplasm, cytosol. Nucleus. Note=Mainly mitochondrial. Also present extramitochondrially: a fraction is present in the cytosol and very small amounts are also detected in the nucleus.
| |
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 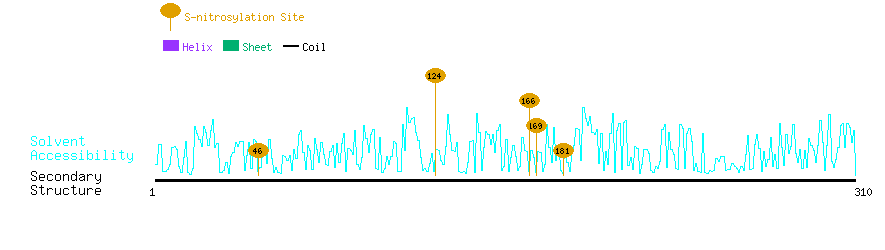 |
|

