Protein Name: NEDD4-like E3 ubiquitin-protein ligase WWP2
|
UniprotKB/SwissProt ID: WWP2_HUMAN (O00308)
Gene Name: WWP2
Organism: Homo sapiens (Human).
Function: E3 ubiquitin-protein ligase which accepts ubiquitin from an E2 ubiquitin-conjugating enzyme in the form of a thioester and then directly transfers the ubiquitin to targeted substrates. Polyubiquitinates POU5F1 by 'Lys-63'-linked conjugation and promotes it to proteasomal degradation; in embryonic stem cells (ESCs) the ubiquitination is proposed to regulate POU5F1 protein level. Ubiquitinates EGR2 and promotes it to proteasomal degradation; in T-cells the ubiquitination inhibits activation- induced cell death. Ubiquitinates SLC11A2; the ubiquitination is enhanced by presence of NDFIP1 and NDFIP2. Ubiquitinates RPB1 and promotes it to proteasomal degradation.
Other Modifications: View all modification sites in dbPTM
Protein Subcellular Localization: Nucleus.
| |
|
Network with metabolic pathway:
|
|
Graphical Visualization of S-nitrosylation Sites:
|
| Overview of Protein S-nitrosylation Sites with Functional and Structural Information | 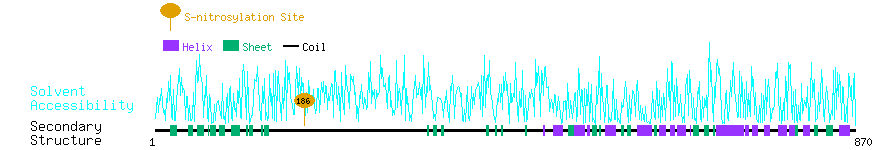 |
|

